|
Here, fastdist is about 27x faster than. cdist ( a, b, "cosine" ) # 689 ms ± 10.3 ms per loop (mean ± std. of 7 runs, 10 loops each) % timeit distance. cosine, "cosine" ) # 25.4 ms ± 1.36 ms per loop (mean ± std. matrix_to_matrix_distance ( a, b, fastdist. rand ( 2500, 1000 ) # i complied the matrix_to_matrix function once before this so it's already in machine code % timeit fastdist. For example: from fastdist import fastdist import numpy as np from scipy.spatial import distance a, b = np. This difference only gets largerĪs the matrices get bigger and when we compile the fastdist function once before running it. In this example, fastdist is about 7x faster than. cdist ( a, b, "cosine" ) # 57.9 ms ± 4.43 ms per loop (mean ± std. of 7 runs, 100 loops each) # note this high stdev is because of the first run taking longer to compile % timeit - n 100 distance. cosine, "cosine" ) # 8.97 ms ± 11.2 ms per loop (mean ± std. Here are some examples comparing the speed of fastdist to : from fastdist import fastdist import numpy as np from scipy.spatial import distance a, b = np. The first runtime includes the compile time. So, the first time you call a function will be slower than the following times, as 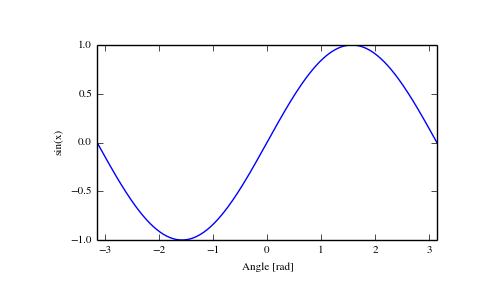
Note that numba - the primary package fastdist uses - compiles the function to machine code the first Matrix/matrix, and pairwise matrix calculations. Notably, cosine similarity is much faster, as are the vector/matrix, Though almost all functions will show a speed improvement in fastdist, certain functions will haveĪn especially large improvement. euclidean, "euclidean", return_matrix = False ) # returns an array of shape (10 choose 2, 1) # to return a matrix with entry (i, j) as the distance between row i and j # set return_matrix=True, in which case this will return a (10, 10) array Speedįastdist is significantly faster than in most cases. euclidean, "euclidean" ) # returns an array of shape (25, 50)įinally, to calculate the pairwise distances between the rows of a matrix, use matrix_pairwise_distance: from fastdist import fastdist import numpy as np a = np. To calculate the distance between the rows of 2 matrices, use matrix_to_matrix_distance: from fastdist import fastdist import numpy as np a = np. euclidean, "euclidean" ) # returns an array of shape (50,) vector_to_matrix_distance ( u, m, fastdist. To calculate the distance between a vector and each row of a matrix, use vector_to_matrix_distance: from fastdist import fastdist import numpy as np u = np. confusion_matrix ( y_true, y_pred )įor calculating distances involving matrices, fastdist has a few different functions instead of scipy's cdist and pdist. So, for example, to create a confusion matrix from two discrete vectors, run: from fastdist import fastdist import numpy as np y_true = np. However, the other functions are the same as trics. Notably, most of the ROC-based functions are not (yet) available in fastdist. The same is true for most trics functions, though not all functions in trics are implemented in fastdist. So, for example, to calculate the Euclidean distance betweenĢ vectors, run: from fastdist import fastdist import numpy as np u = np. pip install fastdistįor calculating the distance between 2 vectors, fastdist uses the same function callsĪs. Use the package manager pip to install fastdist.
0 Comments
Leave a Reply. |
 RSS Feed
RSS Feed
